26 Cytokines
Classification of Cytokines
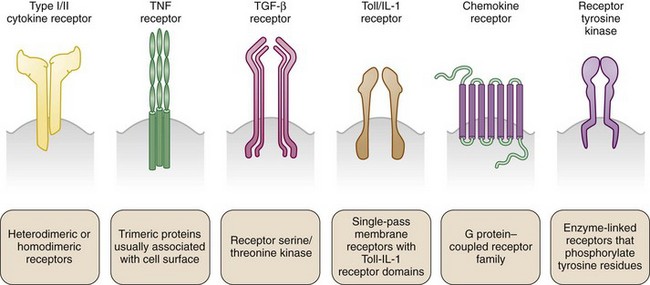
In the absence of a unified classification system, cytokines are variously identified by numeric order of discovery (currently interleukin [IL]-1 through IL-37); by a given functional activity (e.g., tumor necrosis factor [TNF], granulocyte colony-stimulating factor); by kinetic or functional role in inflammatory responses (early or late, innate or adaptive, proinflammatory or anti-inflammatory) by primary cell of origin (monokine = monocyte derivation; lymphokine = lymphocyte derivation); and, more recently, by structural homologies shared with related molecules. Superfamilies of cytokines share sequence similarity and exhibit homology and some promiscuity in their reciprocal receptor systems (Figure 26-1). They do not exhibit functional similarity. Cytokine superfamilies also contain important regulatory cell membrane receptor-ligand pairs, reflecting evolutionary pressures that use common structural motifs in diverse immune functions in higher mammals. The TNF/TNF receptor superfamily1 contains immunoregulatory cytokines including TNF, lymphotoxins, and cellular ligands such as CD40L, which mediates B cell and T cell activation, and FasL (CD95), which promotes apoptosis. Similarly, the IL-1/IL-1 receptor superfamily2 contains cytokines including IL-1β, IL-1α, IL-receptor antagonist, IL-18, IL-33, IL-36 (α,β,γ), IL-36 receptor antagonist, and IL-37 (α,β), which mediate physiologic and host-defense function, but this family also includes the Toll-like receptors, a series of mammalian pattern-recognition molecules with a crucial role in recognition of microbial species early in innate responses.
Assessing Cytokine Function in Vitro And in Vivo
This general approach has been crucial in rheumatic disease research. Studies in which cytokine addition and neutralization occur in synovial tissue explants or disaggregated cell populations, chondrocyte explants, bone culture models, skin, and renal tissue explants and cell lines have been informative. Ex vivo methodologies now include intracellular fluorescence activated cell sorter methods, confocal and laser scanning microscopy, and quantitative histologic evaluation using automated image analysis. Such modalities, particularly when employed in human therapeutic cytokine neutralization studies in which inflammatory tissues are obtained throughout therapeutic interventions, advance the understanding of basic and pathogenetic cytokine function. Analysis of synovial biopsy specimens obtained before and after infliximab, adalimumab, abatacept, rituximab, IL-1Ra, IL-10, and interferon (IFN)-β administration in rheumatoid arthritis provides the strongest evidence for the success of this approach.3,4
Cytokine Receptors
Cytokine receptors exist in structurally related superfamilies and comprise high-affinity molecular signaling complexes that assist cytokine-mediated communication (see Figure 26-1). Such complexes often include heterodimeric or heterotrimeric structures that use unique, cytokine-specific recognition receptors together with common receptor chains shared across a cytokine superfamily. Examples include the use of the common γ chain receptor by IL-2, IL-4, IL-7, IL-9, IL-15, IL-21, and glycoprotein 130 (gp130) by members of the IL-6 family.5,6 Alternatively, distinct receptors may use shared signaling domains. Homologous death domains are found in many TNF-receptor family members. Similarly, the IL-1 signaling domain is common to not only IL-1R but also other IL-1R superfamily members including IL-18R, IL-33R, and the Toll-like receptors.2 Signaling pathways dependent on these are discussed in detail elsewhere. It has been recognized more recently that unrelated cytokine receptor systems exhibit close cross-communication on the cell membrane, allowing a cell to integrate a variety of external stimuli to optimize signaling pathways and the cellular response in real time in a changing environment. Although best elucidated in the epidermal growth factor receptor system, this also has been identified for members of the common γ chain signaling family.
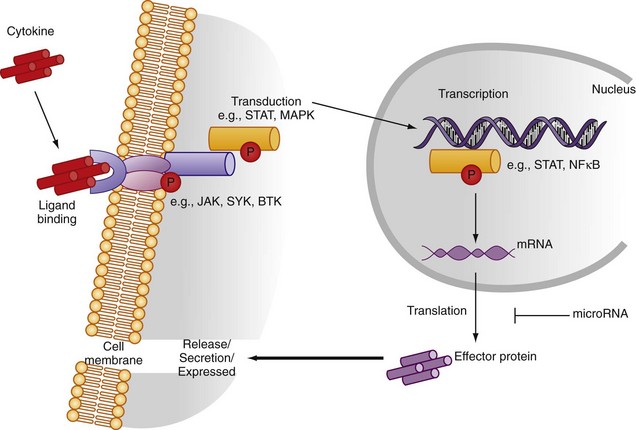
Cytokine receptors can operate via several mechanisms. Membrane receptors, with intracellular signaling domains intact, can transmit signals to the target cell nucleus after soluble cytokine binding and promote effector function (Figure 26-2). Membrane receptors may bind cell membrane cytokines assisting cross-talk between adjacent cells. Membrane-bound and soluble cytokines may promote distinct receptor function. Useful exemplars exist relevant to the rheumatic diseases. Thus TNF binds TNF-RI and TNF-RII with similar affinity, but it has a slower rate of dissociation from TNF-RI. Soluble TNF may dissociate rapidly from TNF-RII to bind TNF-RI, promoting preferential signaling by the latter (ligand passing).1 In contrast, during cell-cell contact, stable TNF/TNF-RI and TNF/TNF-RII complexes form, allowing for differential signaling contribution by TNF-RI and TNF-RII.
Cytokine receptor/cytokine complexes also may operate in trans, whereby component parts of the ligand-receptor complex are derived from adjacent cells. IL-15/IL-15Rα complexed on one cell may bind IL-15Rβ/γ on another.7 Receptors also exist in soluble form, derived either from alternative mRNA processing to generate receptor-lacking transmembrane or intracellular domains or from enzymatic cleavage of receptor from the cell surface (e.g., sTNF-R, sIL-1R1). Soluble receptors may act to antagonize cytokine function, regulating responses. Soluble receptors also may preform complexes with cytokine to promote subsequent ligand-receptor assembly on the target cell membrane and enhance function. Soluble receptors can deliver cytokine to the cell membrane via ligand passing. IL-6 provides a particularly important example given its core role in a range of rheumatic disorders. IL-6 binds to a heterodimeric receptor (IL-6R and gp130) and provokes cell activation thereby via conventional signal pathways that involve STAT3. Thus IL-6 may activate a cell expressing the combination of IL-6R and gp130 by conventional (cis) signaling. In addition, however, circulating soluble IL-6R may form functional gp130/IL-6R effector complexes on any cell expressing membrane gp130 and by this means confer on circulating IL-6 the ability to exert broad functional effects (trans signaling). Finally, it is now recognized that some cytokines with the capacity to be retained in the membrane may themselves function as signaling molecules (reverse signaling).
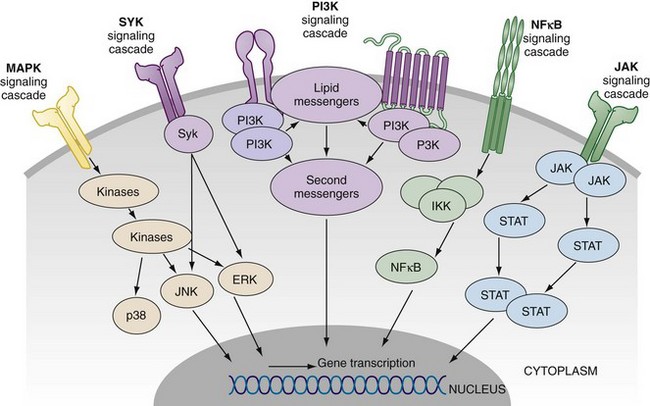
The precise intracellular signal pathways whereby cytokines mediate their effects have now been elucidated and reveal a multiplicity of potential therapeutic targets (Figure 26-3). This is exemplified in the role of the Janus kinases (JAK) and their downstream transcriptional factors (e.g., STATs) in mediating signals serving those cytokines that bind to common gamma chain, or to gp130 receptor. Similarly the mitogen-activated protein (MAP) kinases mediate signals that integrate cell responses to stress, cytokines, and proliferation factors. Spleen tyrosine kinase (Syk) is an additional kinase that mediates the effector function of immunoglobulin Fc receptor and the B cell receptor and interacts with many downstream cytokine-mediated signal pathways. Finally, the NFκB pathway is central to the function of the TNF receptor and to those pathways that mediate signals of Toll-like receptors and IL-1 superfamily members. There is considerable interest in targeting upstream members of the MAP kinase family, JAKs, and Syk, with the latter two demonstrating efficacy in clinical trials focused on rheumatoid arthritis. One key question will be whether the broader cytokine-regulating effects of signal transduction inhibitors will have an acceptable safety profile compared with biologics that target individual cytokines or receptors.
Regulation of Cytokine Expression
Cytokines are synthesized in the Golgi apparatus and may traffic through the endoplasmic reticulum to be released as soluble mediators, or they may remain membrane bound, or they may be processed into cytosolic forms that can traffic intracellularly, even returning to the nucleus, where they can act as transcriptional regulators. Cytokines mediate autocrine function either through release or membrane expression and immediate receptor ligation on the source cell or intracellularly within the source cell. Alternatively, cytokines operate in a paracrine manner, allowing cellular communication beyond that assisted by local cell-cell contact. The distance and kinetics for effective function may be limited8; however, by numerous factors including physicochemical considerations of the peptide structure itself, extracellular matrix binding (e.g., to heparan sulfate), enzymatic degradation (e.g., serine protease degradation of IL-18), or the presence of soluble receptors (e.g., TNF/soluble TNF-RI and TNF-RII, IL-2/soluble IL-2Rα) or novel cytokine-binding proteins (e.g., IL-18/IL-18 binding protein) in the inflammatory milieu.
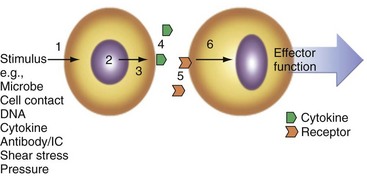
Numerous factors promote cytokine expression in vivo (Figure 26-4) including cell-cell contact, immune complexes/autoantibodies, local complement activation, microbial species and their soluble products (particularly via TLR and nucleotide oligomerization domain [NOD]-like receptors [NLRs]), reactive oxygen and nitrogen intermediates, trauma, shear stress, ischemia, radiation, ultraviolet light, extracellular matrix components, DNA (mammalian or microbial), heat shock proteins, electrolytes (e.g., K+ via P2X7 receptors), and cytokines themselves operating in autocrine loops. Commonly used in vitro stimuli include many of these and chemical entities including phorbol esters, calcium ionophores, lectins (e.g., phytohemagglutinin), and receptor-specific antibodies such as anti-CD3 and anti-CD28 for T cell activation or anti-immunoglobulin and anti-CD40 for B cells.
Cytokine regulation within the cell can be usefully considered at several levels (see Figure 26-2). Transcriptional regulation depends on the recruitment of discrete transcription factors to the cytokine promoter region. Transcription factor binding allows for numerous signal pathways to regulate cytokine expression across a range of stimuli. Several transcription factors (e.g., nuclear factor κB [NFκB], activator protein-1 [AP-1], nuclear factor of activated T cell) are crucial in cytokine production. Inhibition of NFκB activity using either chemical inhibitors or adenoviral delivery of regulatory peptides leads to amelioration of inflammatory synovitis in vivo and in vitro.9 Sequence polymorphism within cytokine promoters offers potential for differential cytokine expression between individuals that could confer selective advantage against infection but could also increase susceptibility to, or progression of, autoimmunity. This is best exemplified in the TNF and IL-1 promoters.10,11 Single nucleotide polymorphisms in the TNF promoter region (e.g., −308) are associated with altered TNF release on leukocyte stimulation in vitro. Similarly, homozygotes for the A2 allele at +3954 in the IL-1β gene produce more IL-1β with lipopolysaccharide stimulation. Polymorphisms also exist in the IL-1Ra gene, rendering the functional significance of individual single nucleotide polymorphisms on IL-1 protein release difficult to interpret. In general, the net effect of haplotypes may be more important at the functional level or only play a role in the context of networks where multiple minor polymorphisms can synergize, particularly when their relevance to disease entities is considered.
Posttranscriptional regulation is important in determining longevity of cytokine expression. This regulation may operate by promoting translational initiation, mRNA stability, and polyadenylation. AU-rich elements (AREs) within the 52 or 32 untranslated regions (UTRs) of cytokine mRNA are crucial for stability; 32 UTR AREs downregulate TNF expression such that transgenic knockin mice that lack TNF AREs develop spontaneous inflammatory arthritis and bowel disease.12 Regulatory proteins bind AREs to mediate such effects. HuR and AUF1 exert opposing effects, stabilizing and destabilizing ARE-containing transcripts.13 TIA-1 and TIAR have been identified as RNA recognition motif family members14 that function as translational silencers. Macrophages from TIA-1-deficient macrophages produce excess TNF, whereas TIA-1-deficient lymphocytes exhibit normal TNF release, suggesting distinctions in mRNA regulation in discrete cell types.15 Alternatively, cytokines may generate stable mRNA a priori to assist subsequent rapid response in tissues. IL-15 mRNA 52 UTR contains 12 AUG triplets that significantly reduce the efficiency of IL-15 translation. Deletion of this sequence permits IL-15 secretion. IL-15 mRNA can produce a 48 amino acid signal peptide that allows IL-15 release and a shorter 21 amino acid signal peptide that targets intracellular distribution. IL-15 forms thus generated exhibit discrete functions.16
Posttranslational regulation also modulates cytokine expression via several mechanisms. Patterns of glycosylation are important for cytokine function and may regulate intracellular trafficking.16 Modified leader sequences can alter intracellular trafficking of cytokines. Some cytokines are translated without functional leader sequences. Their secretion depends on nonconventional secretory pathways that are poorly understood. IL-1β employs, among other pathways, a purine receptor–dependent pathway (P2X7) for cellular release.17 Enzymatic activation of cytokines is common, whereby nonfunctional promolecules are cleaved to generate functional subunits. Examples include the cleavage by caspase 1 of pro-IL-1β to generate active IL-1β and, similarly, of pro-IL-18 to generate an active 18-kD species.18 This is an organized process both sequentially (in time) and by orientation within the cell (in space). IL-1 processing occurs in a protein complex within the cytosol termed the inflammasome. The latter has recently attracted considerable interest as a therapeutic target in conditions such as crystal-induced arthritis and in diseases that arise from mutations in certain inflammasome genes (e.g., cryopyrin) (see Chapters 18, 94, and 97).
Alternative processing pathways for cytokines include the serine proteases, proteinase 3 and elastase, and adamalysin family members. Enzyme cleavage pathways operate within and outside cells, providing for extracellular cytokine activation. Similarly, cell membrane enzymes serve to cleave membrane-expressed cytokine. Members of the adamalysin family regulate TNF release; TNF-converting enzyme cleaves and mediates the release of TNF and its receptors.19 Extensive molecular machinery exists to regulate tightly not only the production and stability of cytokine mRNA but also its translation and cellular expression and distribution. At each level, opportunities exist for intervention and therapeutic cytokine modulation.